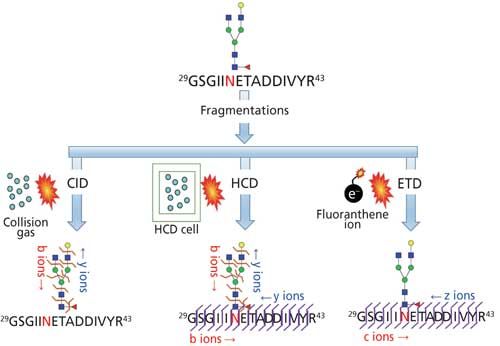
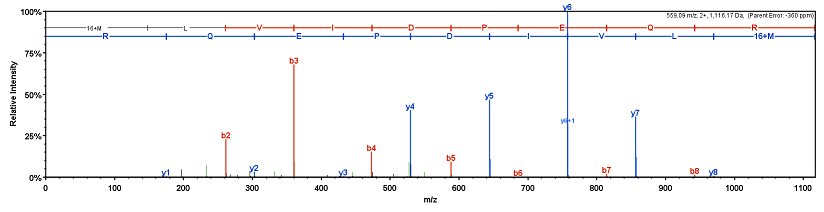
Higher Energy Collision Dissociation (HCD) Product Ion-Triggered Electron Transfer Dissociation (ETD) Mass Spectrometry for the Analysis of N-Linked Glycoproteins | Journal of Proteome Research

Expediting SRM Assay Development for Large-Scale Targeted Proteomics Experiments | Journal of Proteome Research

Effectiveness of CID, HCD, and ETD with FT MS/MS for Degradomic-Peptidomic Analysis: Comparison of Peptide Identification Methods | Journal of Proteome Research
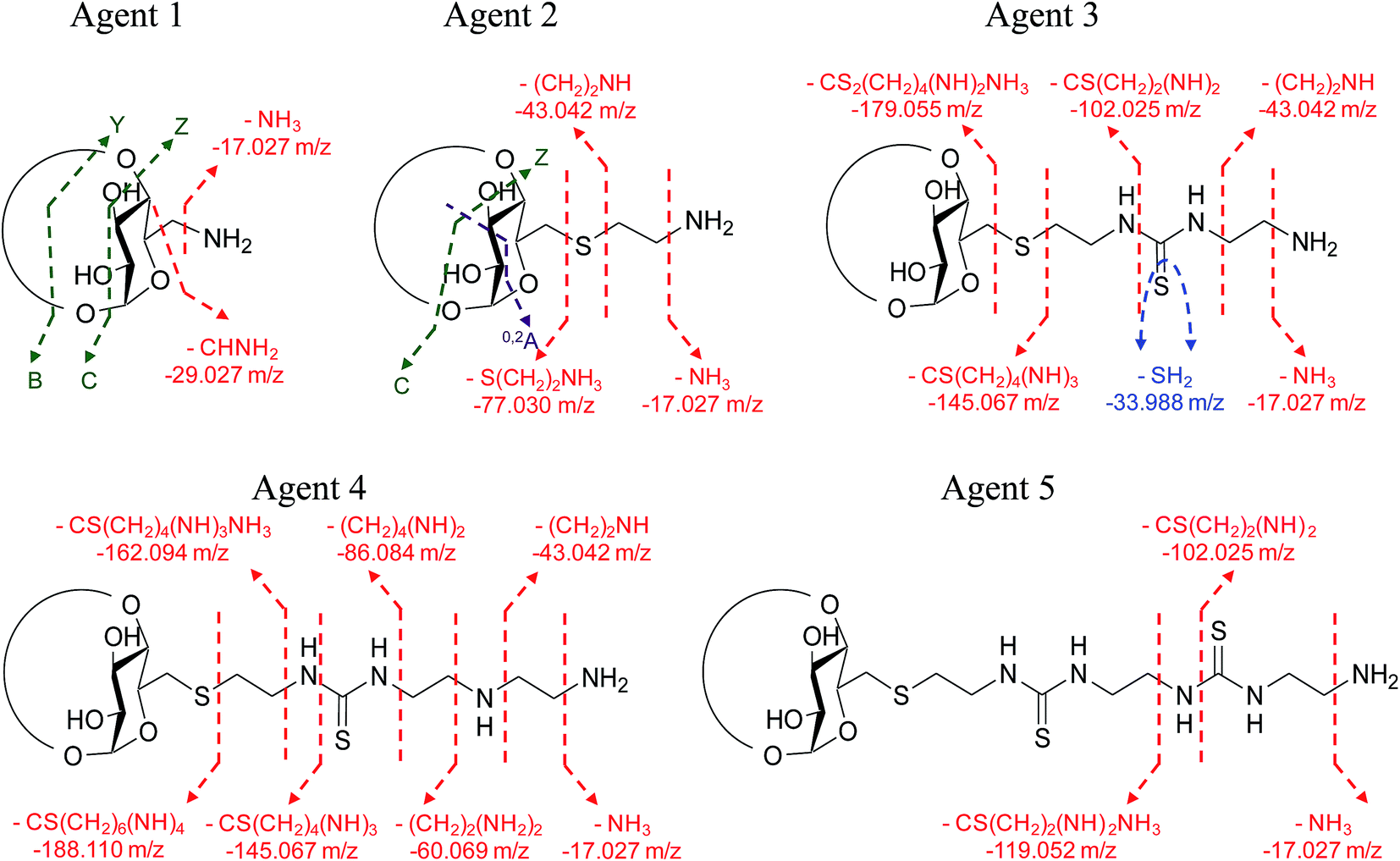
Deciphering of polycationic carbohydrate based non-viral gene delivery agents by ESI-LTQ-Orbitrap using CID/HCD pairwise tandem mass spectrometry - RSC Advances (RSC Publishing) DOI:10.1039/C6RA14508F
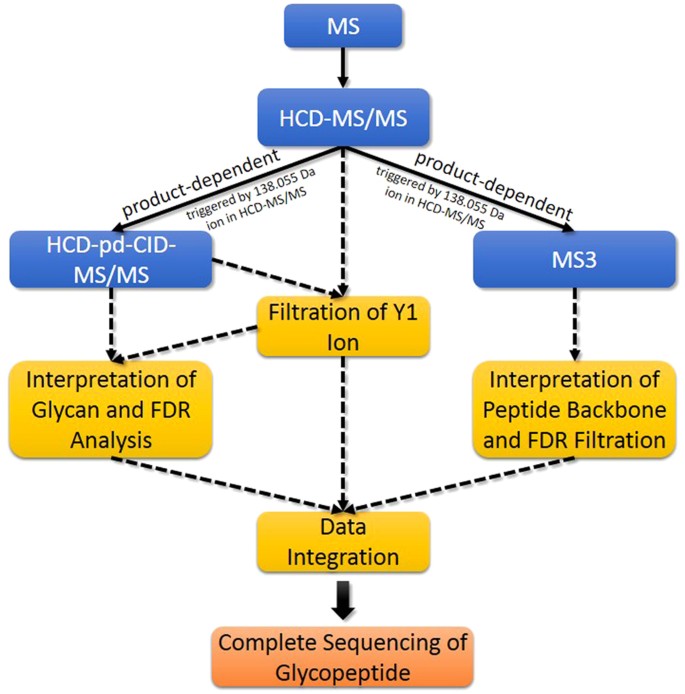
pGlyco: a pipeline for the identification of intact N-glycopeptides by using HCD- and CID-MS/MS and MS3 | Scientific Reports

CIDer: A Statistical Framework for Interpreting Differences in CID and HCD Fragmentation | Journal of Proteome Research
Performance Investigation of Proteomic Identification by HCD/CID Fragmentations in Combination with High/Low-Resolution Detectors on a Tribrid, High-Field Orbitrap Instrument | PLOS ONE

2. A table describing the ion activation methods CID, HCD, and ETD. The... | Download Scientific Diagram
Performance Investigation of Proteomic Identification by HCD/CID Fragmentations in Combination with High/Low-Resolution Detectors on a Tribrid, High-Field Orbitrap Instrument | PLOS ONE

Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics - ScienceDirect

Characterization of collision-induced dissociation of deprotonated peptides of 4–16 amino acids using high-resolution mass spectrometry - ScienceDirect
![PDF] Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics* | Semantic Scholar PDF] Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics* | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/d76885d896eb57c5dcb781600ed39724f4c46016/7-Figure4-1.png)
PDF] Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics* | Semantic Scholar

Finding and using diagnostic ions in collision induced crosslinked peptide fragmentation spectra - ScienceDirect
Performance Investigation of Proteomic Identification by HCD/CID Fragmentations in Combination with High/Low-Resolution Detectors on a Tribrid, High-Field Orbitrap Instrument | PLOS ONE