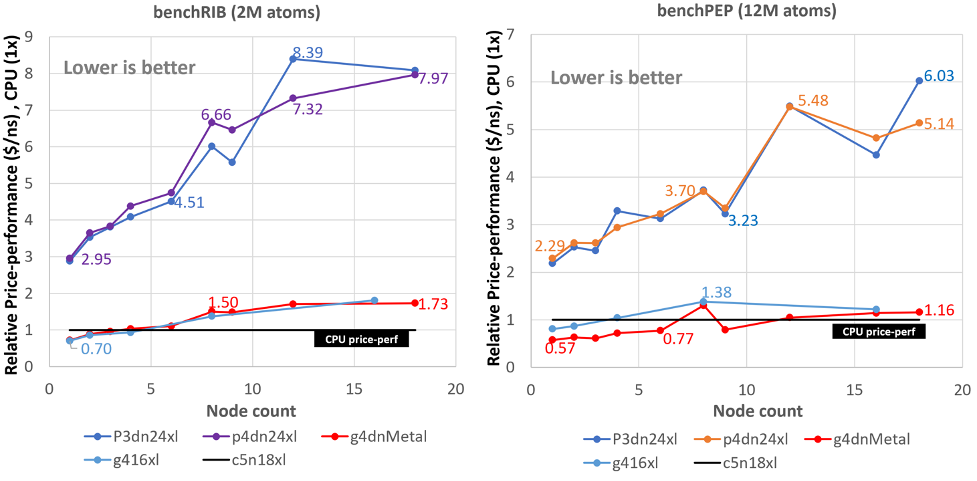
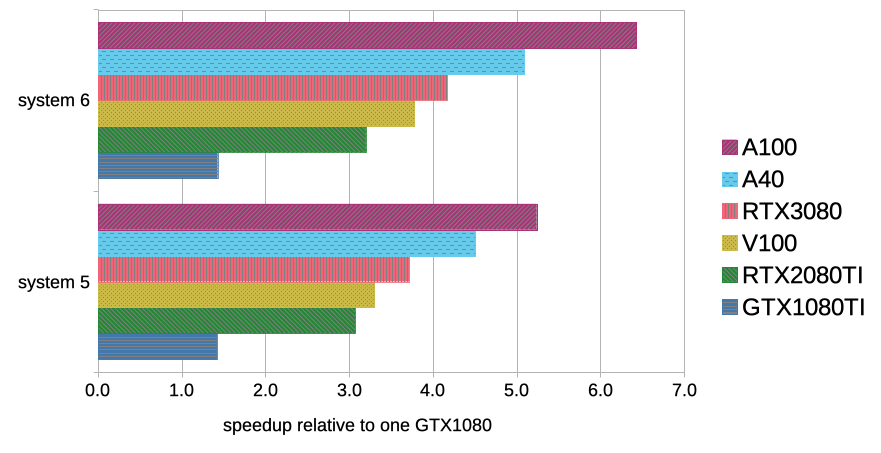
Best bang for your buck: GPU nodes for GROMACS biomolecular simulations - Kutzner - 2015 - Journal of Computational Chemistry - Wiley Online Library

A GPU-Accelerated Fast Multipole Method for GROMACS: Performance and Accuracy | Journal of Chemical Theory and Computation
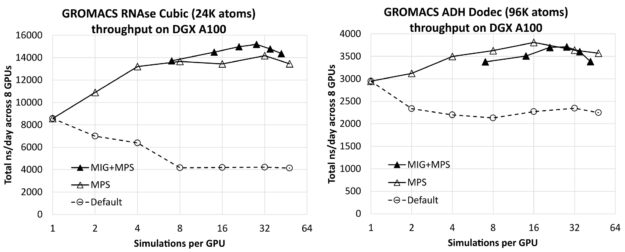
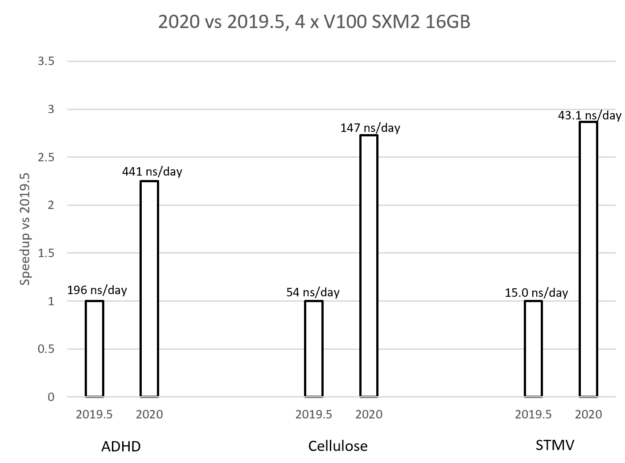
Maximizing GROMACS Throughput with Multiple Simulations per GPU Using MPS and MIG | NVIDIA Technical Blog

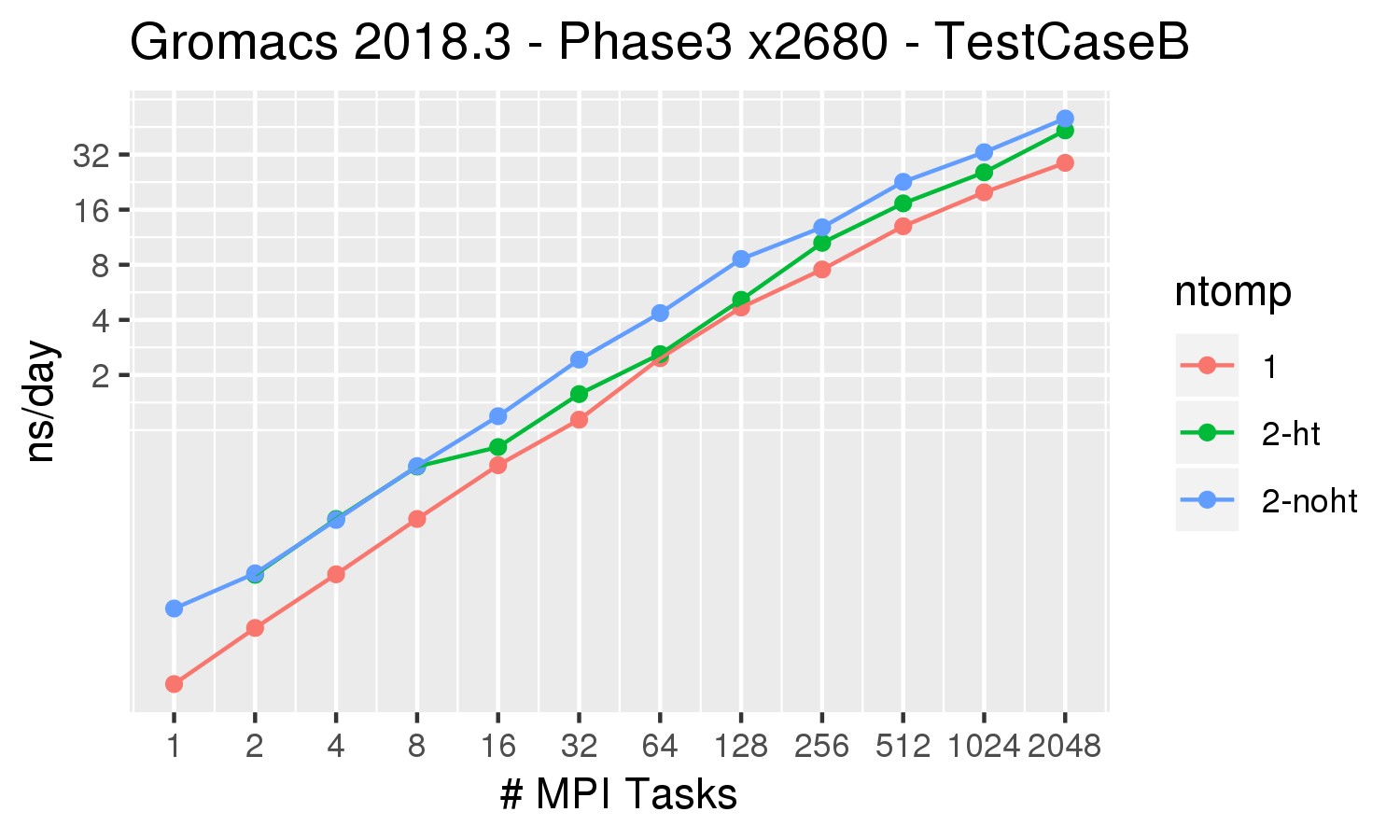
GROMACS scaling on HYDRA (Max Planck Computing Centre), reproduced from... | Download Scientific Diagram

Heterogeneous Parallelization and Acceleration of Molecular Dynamics Simulations in GROMACS | DeepAI

A GPU-Accelerated Fast Multipole Method for GROMACS: Performance and Accuracy | Journal of Chemical Theory and Computation

A GPU-Accelerated Fast Multipole Method for GROMACS: Performance and Accuracy | Journal of Chemical Theory and Computation

GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers - ScienceDirect