PureseqTM: efficient and accurate prediction of transmembrane topology from amino acid sequence only | bioRxiv

Improved sequence-based prediction of interaction sites in α-helical transmembrane proteins by deep learning - ScienceDirect

MemBrain: An Easy-to-Use Online Webserver for Transmembrane Protein Structure Prediction | SpringerLink
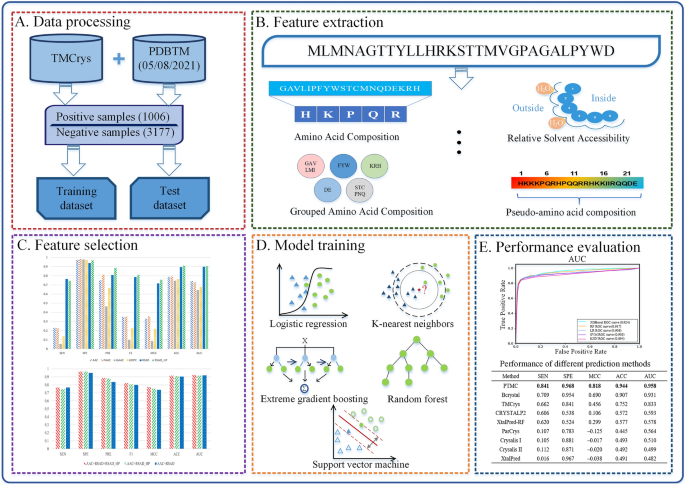
Membrane contact probability: An essential and predictive character for the structural and functional studies of membrane proteins | PLOS Computational Biology

IJMS | Free Full-Text | RLPredictiOme, a Machine Learning-Derived Method for High-Throughput Prediction of Plant Receptor-like Proteins, Reveals Novel Classes of Transmembrane Receptors

BetAware-Deep: An Accurate Web Server for Discrimination and Topology Prediction of Prokaryotic Transmembrane β-barrel Proteins - ScienceDirect
Membrane contact probability: An essential and predictive character for the structural and functional studies of membrane proteins | PLOS Computational Biology

PureseqTM: efficient and accurate prediction of transmembrane topology from amino acid sequence only | bioRxiv
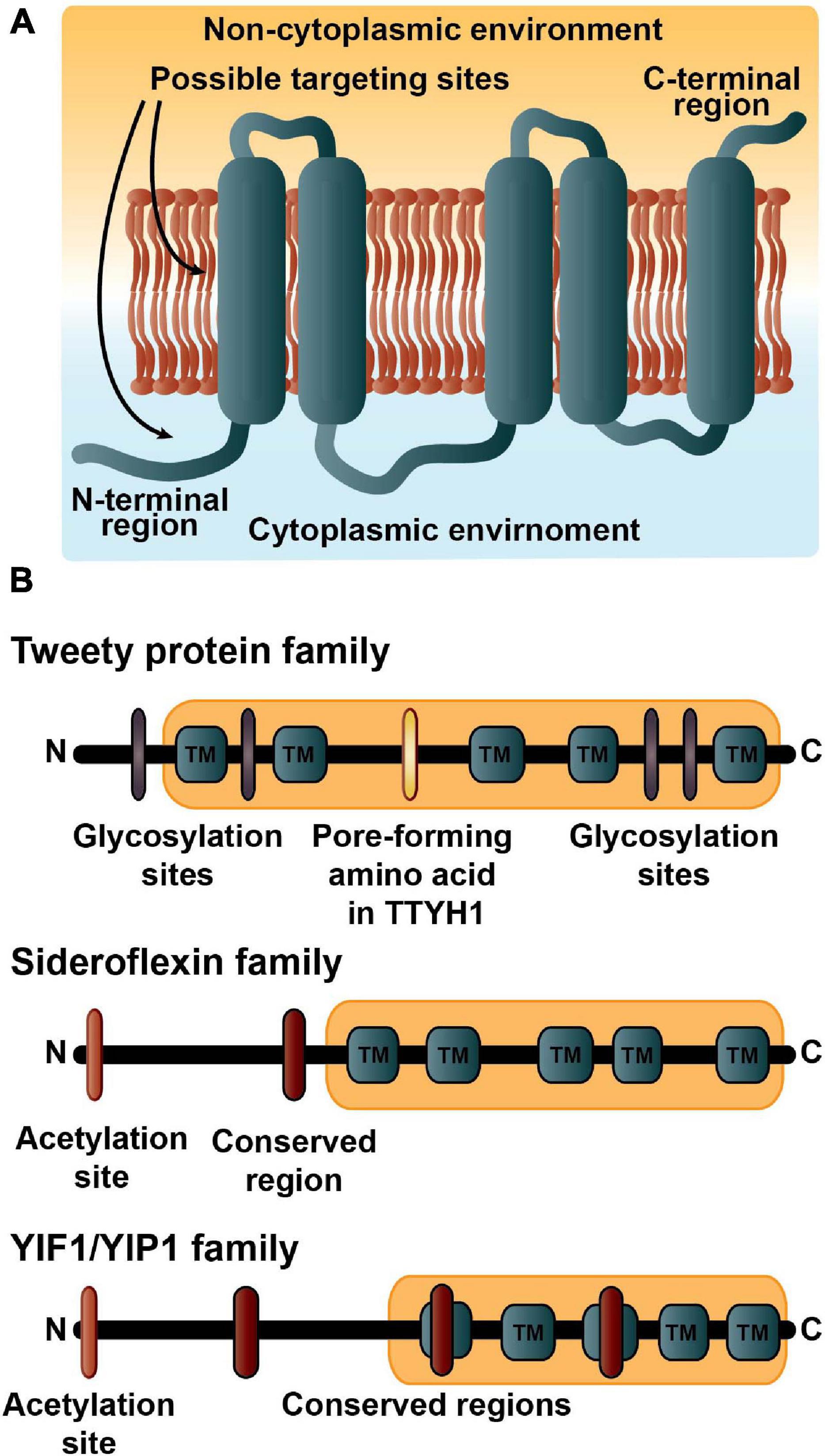
Frontiers | Characterization of Five Transmembrane Proteins: With Focus on the Tweety, Sideroflexin, and YIP1 Domain Families

MPTherm-pred: Analysis and Prediction of Thermal Stability Changes upon Mutations in Transmembrane Proteins - ScienceDirect
![PDF] A Hidden Markov Model for Predicting Transmembrane Helices in Protein Sequences | Semantic Scholar PDF] A Hidden Markov Model for Predicting Transmembrane Helices in Protein Sequences | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/023499b96536b1a0dcb04a48578c5ee2533237bc/3-Figure1-1.png)